How to Read 2 Number Pairs Into an Array of Structures in C#
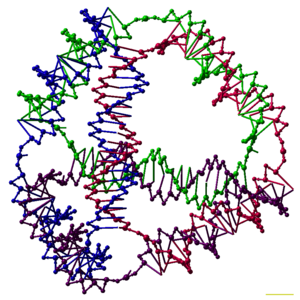
DNA nanotechnology involves forming bogus, designed nanostructures out of nucleic acids, such as this Deoxyribonucleic acid tetrahedron.[1] Each border of the tetrahedron is a 20 base of operations pair Dna double helix, and each vertex is a three-arm junction. The iv Deoxyribonucleic acid strands that form the iv tetrahedral faces are color-coded.
Deoxyribonucleic acid nanotechnology is the design and manufacture of artificial nucleic acid structures for technological uses. In this field, nucleic acids are used as not-biological engineering materials for nanotechnology rather than every bit the carriers of genetic information in living cells. Researchers in the field have created static structures such as two- and iii-dimensional crystal lattices, nanotubes, polyhedra, and arbitrary shapes, and functional devices such as molecular machines and DNA computers. The field is start to be used as a tool to solve basic science problems in structural biological science and biophysics, including applications in X-ray crystallography and nuclear magnetic resonance spectroscopy of proteins to determine structures. Potential applications in molecular calibration electronics and nanomedicine are as well being investigated.
The conceptual foundation for Deoxyribonucleic acid nanotechnology was showtime laid out past Nadrian Seeman in the early on 1980s, and the field began to attract widespread involvement in the mid-2000s. This use of nucleic acids is enabled by their strict base pairing rules, which cause but portions of strands with complementary base of operations sequences to demark together to class stiff, rigid double helix structures. This allows for the rational design of base sequences that will selectively assemble to form complex target structures with precisely controlled nanoscale features. Several assembly methods are used to make these structures, including tile-based structures that assemble from smaller structures, folding structures using the DNA origami method, and dynamically reconfigurable structures using strand displacement methods. The field's proper name specifically references Deoxyribonucleic acid, but the aforementioned principles have been used with other types of nucleic acids as well, leading to the occasional utilise of the alternative name nucleic acid nanotechnology.
Fundamental concepts [edit]
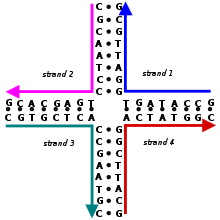
These four strands associate into a DNA 4-arm junction because this structure maximizes the number of right base pairs, with A matched to T and C matched to G.[2] [three] Run across this prototype for a more realistic model of the 4-arm junction showing its tertiary structure.

This double-crossover (DX) supramolecular complex consists of five DNA unmarried strands that class two double-helical domains, on the top and the bottom in this image. There are two crossover points where the strands cross from one domain into the other.[ii]
Properties of nucleic acids [edit]
Nanotechnology is oft defined as the written report of materials and devices with features on a scale below 100 nanometers. Dna nanotechnology, specifically, is an case of lesser-up molecular self-assembly, in which molecular components spontaneously organize into stable structures; the item form of these structures is induced by the concrete and chemical properties of the components selected by the designers.[four] In Dna nanotechnology, the component materials are strands of nucleic acids such equally Deoxyribonucleic acid; these strands are oftentimes constructed and are almost e'er used exterior the context of a living jail cell. DNA is well-suited to nanoscale construction considering the binding betwixt two nucleic acid strands depends on simple base pairing rules which are well understood, and form the specific nanoscale structure of the nucleic acrid double helix. These qualities make the assembly of nucleic acid structures easy to control through nucleic acid design. This property is absent in other materials used in nanotechnology, including proteins, for which protein design is very difficult, and nanoparticles, which lack the capability for specific assembly on their ain.[five]
The construction of a nucleic acid molecule consists of a sequence of nucleotides distinguished by which nucleobase they comprise. In DNA, the four bases present are adenine (A), cytosine (C), guanine (G), and thymine (T). Nucleic acids accept the property that two molecules will only bind to each other to grade a double helix if the 2 sequences are complementary, significant that they form matching sequences of base pairs, with A but binding to T, and C only to Yard.[v] [6] Because the germination of correctly matched base pairs is energetically favorable, nucleic acid strands are expected in well-nigh cases to bind to each other in the conformation that maximizes the number of correctly paired bases. The sequences of bases in a system of strands thus determine the design of binding and the overall structure in an hands controllable way. In Dna nanotechnology, the base of operations sequences of strands are rationally designed by researchers so that the base pairing interactions cause the strands to assemble in the desired conformation.[iii] [v] While DNA is the dominant material used, structures incorporating other nucleic acids such equally RNA and peptide nucleic acid (PNA) have also been constructed.[7] [eight]
Subfields [edit]
Deoxyribonucleic acid nanotechnology is sometimes divided into two overlapping subfields: structural Deoxyribonucleic acid nanotechnology and dynamic DNA nanotechnology. Structural DNA nanotechnology, sometimes abbreviated as SDN, focuses on synthesizing and characterizing nucleic acid complexes and materials that assemble into a static, equilibrium stop country. On the other mitt, dynamic DNA nanotechnology focuses on complexes with useful non-equilibrium behavior such as the ability to reconfigure based on a chemical or concrete stimulus. Some complexes, such as nucleic acid nanomechanical devices, combine features of both the structural and dynamic subfields.[nine] [10]
The complexes constructed in structural DNA nanotechnology employ topologically branched nucleic acid structures containing junctions. (In dissimilarity, most biological Dna exists equally an unbranched double helix.) One of the simplest branched structures is a four-arm junction that consists of four individual Dna strands, portions of which are complementary in a specific blueprint. Unlike in natural Holliday junctions, each arm in the artificial immobile four-arm junction has a different base sequence, causing the junction point to be fixed at a certain position. Multiple junctions can be combined in the same circuitous, such as in the widely used double-crossover (DX) structural motif, which contains two parallel double helical domains with individual strands crossing between the domains at two crossover points. Each crossover point is, topologically, a four-arm junction, but is constrained to one orientation, in contrast to the flexible single four-arm junction, providing a rigidity that makes the DX motif suitable equally a structural building block for larger Dna complexes.[3] [5]
Dynamic DNA nanotechnology uses a mechanism called toehold-mediated strand displacement to permit the nucleic acrid complexes to reconfigure in response to the addition of a new nucleic acid strand. In this reaction, the incoming strand binds to a unmarried-stranded toehold region of a double-stranded complex, then displaces one of the strands spring in the original complex through a branch migration process. The overall effect is that one of the strands in the circuitous is replaced with another i.[9] In addition, reconfigurable structures and devices tin can be made using functional nucleic acids such as deoxyribozymes and ribozymes, which can perform chemical reactions, and aptamers, which can bind to specific proteins or small-scale molecules.[11]
Structural DNA nanotechnology [edit]
Structural Dna nanotechnology, sometimes abbreviated as SDN, focuses on synthesizing and characterizing nucleic acid complexes and materials where the assembly has a static, equilibrium endpoint. The nucleic acid double helix has a robust, defined three-dimensional geometry that makes it possible to simulate,[12] predict and design the structures of more complicated nucleic acrid complexes. Many such structures have been created, including two- and three-dimensional structures, and periodic, aperiodic, and discrete structures.[10]
Extended lattices [edit]
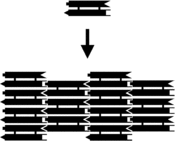

The assembly of a DX assortment. Left, schematic diagram. Each bar represents a double-helical domain of Deoxyribonucleic acid, with the shapes representing complementary sticky ends. The DX complex at top will combine with other DX complexes into the two-dimensional array shown at bottom.[ii] Right, an atomic force microscopy paradigm of the assembled array. The individual DX tiles are clearly visible within the assembled structure. The field is 150 nm across.
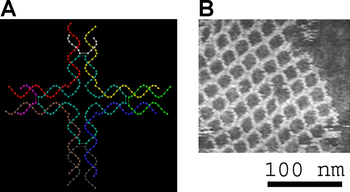

Left, a model of a Deoxyribonucleic acid tile used to brand some other two-dimensional periodic lattice. Correct, an atomic force micrograph of the assembled lattice.[13] [14]
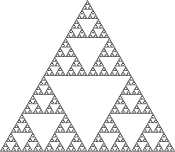


An example of an aperiodic ii-dimensional lattice that assembles into a fractal pattern. Left, the Sierpinski gasket fractal. Right, Deoxyribonucleic acid arrays that display a representation of the Sierpinski gasket on their surfaces[15]
Small nucleic acid complexes can be equipped with gluey ends and combined into larger two-dimensional periodic lattices containing a specific tessellated pattern of the individual molecular tiles.[10] The earliest example of this used double-crossover (DX) complexes as the bones tiles, each containing four sticky ends designed with sequences that caused the DX units to combine into periodic two-dimensional flat sheets that are essentially rigid two-dimensional crystals of Deoxyribonucleic acid.[xvi] [17] Two-dimensional arrays accept been made from other motifs too, including the Holliday junction rhombus lattice,[18] and various DX-based arrays making use of a double-cohesion scheme.[19] [xx] The top two images at right prove examples of tile-based periodic lattices.
Two-dimensional arrays tin can exist made to showroom aperiodic structures whose assembly implements a specific algorithm, exhibiting one form of Deoxyribonucleic acid calculating.[21] The DX tiles tin have their gluey terminate sequences chosen so that they human activity as Wang tiles, assuasive them to perform ciphering. A DX assortment whose assembly encodes an XOR performance has been demonstrated; this allows the DNA assortment to implement a cellular automaton that generates a fractal known as the Sierpinski gasket. The third epitome at right shows this type of array.[15] Another system has the function of a binary counter, displaying a representation of increasing binary numbers equally it grows. These results show that ciphering tin can exist incorporated into the assembly of Dna arrays.[22]
DX arrays have been fabricated to class hollow nanotubes 4–xx nm in diameter, essentially two-dimensional lattices which bend back upon themselves.[23] These Dna nanotubes are somewhat similar in size and shape to carbon nanotubes, and while they lack the electrical conductance of carbon nanotubes, Dna nanotubes are more than easily modified and connected to other structures. Ane of many schemes for constructing Dna nanotubes uses a lattice of curved DX tiles that curls around itself and closes into a tube.[24] In an culling method that allows the circumference to be specified in a simple, modular fashion using single-stranded tiles, the rigidity of the tube is an emergent property.[25]
Forming three-dimensional lattices of Deoxyribonucleic acid was the earliest goal of Deoxyribonucleic acid nanotechnology, but this proved to be ane of the near difficult to realize. Success using a motif based on the concept of tensegrity, a balance between tension and compression forces, was finally reported in 2009.[21] [26]
Discrete structures [edit]
Researchers have synthesized many three-dimensional Deoxyribonucleic acid complexes that each accept the connectivity of a polyhedron, such as a cube or octahedron, meaning that the Dna duplexes trace the edges of a polyhedron with a DNA junction at each vertex.[27] The earliest demonstrations of DNA polyhedra were very piece of work-intensive, requiring multiple ligations and solid-phase synthesis steps to create catenated polyhedra.[28] Subsequent piece of work yielded polyhedra whose synthesis was much easier. These include a DNA octahedron fabricated from a long single strand designed to fold into the right conformation,[29] and a tetrahedron that can be produced from four DNA strands in one step, pictured at the meridian of this article.[1]
Nanostructures of arbitrary, not-regular shapes are commonly made using the Dna origami method. These structures consist of a long, natural virus strand as a "scaffold", which is made to fold into the desired shape by computationally designed short "staple" strands. This method has the advantages of existence easy to blueprint, every bit the base sequence is predetermined by the scaffold strand sequence, and not requiring high strand purity and authentic stoichiometry, as most other DNA nanotechnology methods practise. DNA origami was first demonstrated for ii-dimensional shapes, such as a smiley face, a coarse map of the Western Hemisphere, and the Mona Lisa painting.[27] [30] [31] Solid iii-dimensional structures tin can exist made by using parallel DNA helices arranged in a honeycomb pattern,[32] and structures with two-dimensional faces can be fabricated to fold into a hollow overall three-dimensional shape, alike to a cardboard box. These can be programmed to open and reveal or release a molecular cargo in response to a stimulus, making them potentially useful equally programmable molecular cages.[33] [34]
Templated assembly [edit]
Nucleic acid structures tin be made to comprise molecules other than nucleic acids, sometimes called heteroelements, including proteins, metallic nanoparticles, quantum dots, and fullerenes. This allows the structure of materials and devices with a range of functionalities much greater than is possible with nucleic acids lone. The goal is to use the self-associates of the nucleic acrid structures to template the assembly of the nanoparticles hosted on them, controlling their position and in some cases orientation.[27] [35] Many of these schemes apply a covalent zipper scheme, using oligonucleotides with amide or thiol functional groups every bit a chemical handle to bind the heteroelements. This covalent binding scheme has been used to accommodate gold nanoparticles on a DX-based array,[36] and to adapt streptavidin protein molecules into specific patterns on a DX array.[37] A non-covalent hosting scheme using Dervan polyamides on a DX assortment was used to arrange streptavidin proteins in a specific pattern on a DX array.[38] Carbon nanotubes have been hosted on Deoxyribonucleic acid arrays in a pattern assuasive the assembly to act equally a molecular electronic device, a carbon nanotube field-issue transistor.[39] In improver, there are nucleic acid metallization methods, in which the nucleic acrid is replaced by a metal which assumes the general shape of the original nucleic acid structure,[40] and schemes for using nucleic acid nanostructures as lithography masks, transferring their pattern into a solid surface.[41]
Dynamic Dna nanotechnology [edit]
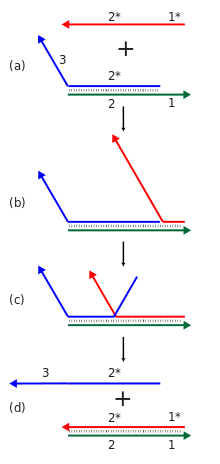
Dynamic Deoxyribonucleic acid nanotechnology ofttimes makes apply of toehold-mediated strand displacement reactions. In this case, the red strand binds to the single stranded toehold region on the green strand (region 1), and and then in a co-operative migration procedure beyond region 2, the bluish strand is displaced and freed from the circuitous. Reactions similar these are used to dynamically reconfigure or assemble nucleic acid nanostructures. In add-on, the blood-red and blue strands can be used equally signals in a molecular logic gate.
Dynamic Deoxyribonucleic acid nanotechnology focuses on forming nucleic acid systems with designed dynamic functionalities related to their overall structures, such every bit computation and mechanical motion. There is some overlap betwixt structural and dynamic Deoxyribonucleic acid nanotechnology, as structures can be formed through annealing and and so reconfigured dynamically, or can be fabricated to grade dynamically in the start identify.[27] [42]
Nanomechanical devices [edit]
DNA complexes take been made that change their conformation upon some stimulus, making them one form of nanorobotics. These structures are initially formed in the aforementioned mode as the static structures made in structural DNA nanotechnology, but are designed so that dynamic reconfiguration is possible after the initial associates.[9] [42] The earliest such device fabricated employ of the transition between the B-Deoxyribonucleic acid and Z-DNA forms to reply to a alter in buffer weather by undergoing a twisting motility.[43] This reliance on buffer weather condition caused all devices to change state at the same fourth dimension. Subsequent systems could alter states based upon the presence of command strands, allowing multiple devices to be independently operated in solution. Some examples of such systems are a "molecular tweezers" design that has an open and a closed land,[44] a device that could switch from a paranemic-crossover (PX) conformation to a double-junction (JX2) conformation, undergoing rotational motion in the procedure,[45] and a ii-dimensional array that could dynamically expand and contract in response to command strands.[46] Structures accept also been made that dynamically open or close, potentially acting as a molecular cage to release or reveal a functional cargo upon opening.[33] [47] [48]
DNA walkers are a class of nucleic acid nanomachines that exhibit directional motion forth a linear rails. A large number of schemes take been demonstrated.[42] Ane strategy is to control the motion of the walker along the track using control strands that demand to be manually added in sequence.[49] [50] Information technology is also possible to control individual steps of a Deoxyribonucleic acid walker by irradiation with light of dissimilar wavelengths.[51] Another approach is to make use of restriction enzymes or deoxyribozymes to carve the strands and cause the walker to movement forward, which has the reward of running autonomously.[52] [53] A later organisation could walk upon a ii-dimensional surface rather than a linear rail, and demonstrated the ability to selectively selection up and movement molecular cargo.[54] In 2018, a catenated Deoxyribonucleic acid that uses rolling circle transcription by an fastened T7 RNA polymerase was shown to walk along a Dna-path, guided by the generated RNA strand.[55] Additionally, a linear walker has been demonstrated that performs Dna-templated synthesis equally the walker advances forth the track, allowing autonomous multistep chemical synthesis directed past the walker.[56] The synthetic DNA walkers' function is similar to that of the proteins dynein and kinesin.[57]
Strand displacement cascades [edit]
Cascades of strand deportation reactions can be used for either computational or structural purposes. An individual strand displacement reaction involves revealing a new sequence in response to the presence of some initiator strand. Many such reactions can be linked into a cascade where the newly revealed output sequence of 1 reaction can initiate another strand displacement reaction elsewhere. This in turn allows for the construction of chemic reaction networks with many components, exhibiting complex computational and information processing abilities. These cascades are made energetically favorable through the germination of new base pairs, and the entropy gain from disassembly reactions. Strand displacement cascades allow isothermal operation of the assembly or computational process, in dissimilarity to traditional nucleic acid assembly's requirement for a thermal annealing pace, where the temperature is raised and so slowly lowered to ensure proper formation of the desired structure. They can likewise support catalytic function of the initiator species, where less than 1 equivalent of the initiator can cause the reaction to go to completion.[9] [58]
Strand displacement complexes can be used to brand molecular logic gates capable of complex computation.[59] Dissimilar traditional electronic computers, which apply electric current as inputs and outputs, molecular computers utilise the concentrations of specific chemic species every bit signals. In the case of nucleic acid strand deportation circuits, the betoken is the presence of nucleic acid strands that are released or consumed by binding and unbinding events to other strands in displacement complexes. This approach has been used to make logic gates such as AND, OR, and NOT gates.[60] More than recently, a four-bit excursion was demonstrated that can compute the square root of the integers 0–15, using a system of gates containing 130 DNA strands.[61]
Another use of strand displacement cascades is to make dynamically assembled structures. These utilise a hairpin construction for the reactants, so that when the input strand binds, the newly revealed sequence is on the same molecule rather than disassembling. This allows new opened hairpins to be added to a growing complex. This arroyo has been used to brand unproblematic structures such as iii- and four-arm junctions and dendrimers.[58]
Applications [edit]
DNA nanotechnology provides i of the few ways to form designed, circuitous structures with precise control over nanoscale features. The field is start to see application to solve basic science problems in structural biology and biophysics. The earliest such awarding envisaged for the field, and one notwithstanding in evolution, is in crystallography, where molecules that are difficult to crystallize in isolation could be arranged within a three-dimensional nucleic acid lattice, allowing determination of their structure. Another application is the utilize of Dna origami rods to supersede liquid crystals in rest dipolar coupling experiments in poly peptide NMR spectroscopy; using Deoxyribonucleic acid origami is advantageous because, unlike liquid crystals, they are tolerant of the detergents needed to suspend membrane proteins in solution. DNA walkers take been used as nanoscale assembly lines to move nanoparticles and direct chemical synthesis. Farther, DNA origami structures have aided in the biophysical studies of enzyme function and protein folding.[10] [62]
Deoxyribonucleic acid nanotechnology is moving toward potential real-world applications. The ability of nucleic acid arrays to arrange other molecules indicates its potential applications in molecular scale electronics. The assembly of a nucleic acrid structure could exist used to template the assembly of molecular electronic elements such as molecular wires, providing a method for nanometer-scale control of the placement and overall architecture of the device analogous to a molecular breadboard.[x] [27] DNA nanotechnology has been compared to the concept of programmable matter because of the coupling of computation to its material properties.[63]
In a written report conducted by a group of scientists from iNANO and CDNA centers in Aarhus University, researchers were able to construct a pocket-size multi-switchable 3D DNA Box Origami. The proposed nanoparticle was characterized by atomic force microscopy (AFM), manual electron microscopy (TEM) and Förster resonance energy transfer (FRET). The constructed box was shown to have a unique reclosing mechanism, which enabled it to repeatedly open and shut in response to a unique fix of Dna or RNA keys. The authors proposed that this "Deoxyribonucleic acid device tin can potentially exist used for a wide range of applications such as controlling the part of single molecules, controlled drug delivery, and molecular calculating."[64]
In that location are potential applications for DNA nanotechnology in nanomedicine, making use of its ability to perform computation in a biocompatible format to make "smart drugs" for targeted drug delivery, every bit well every bit for diagnostic applications. One such system beingness investigated uses a hollow DNA box containing proteins that induce apoptosis, or prison cell expiry, that will only open when in proximity to a cancer cell.[62] [65] There has additionally been interest in expressing these bogus structures in engineered living bacterial cells, nearly likely using the transcribed RNA for the assembly, although it is unknown whether these complex structures are able to efficiently fold or assemble in the cell's cytoplasm. If successful, this could enable directed evolution of nucleic acid nanostructures.[27] Scientists at Oxford Academy reported the self-assembly of four short strands of constructed Deoxyribonucleic acid into a cage which can enter cells and survive for at least 48 hours. The fluorescently labeled DNA tetrahedra were found to remain intact in the laboratory cultured human kidney cells despite the attack by cellular enzymes afterward ii days. This experiment showed the potential of drug delivery inside the living cells using the Dna 'cage'.[66] [67] A DNA tetrahedron was used to deliver RNA Interference (RNAi) in a mouse model, reported a squad of researchers in MIT. Delivery of the interfering RNA for treatment has showed some success using polymer or lipid, only there are limits of safety and imprecise targeting, in addition to short shelf life in the blood stream. The Dna nanostructure created by the team consists of six strands of DNA to form a tetrahedron, with i strand of RNA affixed to each of the half dozen edges. The tetrahedron is further equipped with targeting protein, iii folate molecules, which lead the Deoxyribonucleic acid nanoparticles to the abundant folate receptors found on some tumors. The issue showed that the gene expression targeted by the RNAi, luciferase, dropped past more than half. This written report shows hope in using DNA nanotechnology equally an effective tool to deliver treatment using the emerging RNA Interference technology.[68] [69] The DNA tetrahedron was also used in an effort to overcome the phenomena multidrug resistance. Doxorubicin (DOX) was conjugated with the tetrahedron and was loaded into MCF-7 breast cancer cells that contained the P-glycoprotein drug efflux pump. The results of the experiment showed the DOX was not beingness pumped out and apoptosis of the cancer cells was accomplished. The tetrahedron without DOX was loaded into cells to examination its biocompatibility, and the structure showed no cytotoxicity itself.[seventy] The DNA tetrahedron was besides used as barcode for profiling the subcellular expression and distribution of proteins in cells for diagnostic purposes. The tetrahedral-nanostructured showed enhanced signal due to college labeling efficiency and stability.[71]
Applications for DNA nanotechnology in nanomedicine also focus on mimicking the structure and function of naturally occurring membrane proteins with designed DNA nanostructures. In 2012, Langecker et al. [72] introduced a pore-shaped Deoxyribonucleic acid origami construction that can self-insert into lipid membranes via hydrophobic cholesterol modifications and induce ionic currents across the membrane. This showtime demonstration of a constructed DNA ion channel was followed by a diverseness of pore-inducing designs ranging from a single Dna duplex,[73] to small tile-based structures,[74] [75] [76] [77] [78] and big Deoxyribonucleic acid origami transmembrane porins.[79] Similar to naturally occurring protein ion channels, this ensemble of synthetic Dna-made counterparts thereby spans multiple orders of magnitude in conductance. The report of the membrane-inserting single Deoxyribonucleic acid duplex showed that current must likewise flow on the Dna-lipid interface equally no fundamental channel lumen is present in the blueprint that lets ions pass beyond the lipid bilayer. This indicated that the Deoxyribonucleic acid-induced lipid pore has a toroidal shape, rather than cylindrical, every bit lipid headgroups reorient to face towards the membrane-inserted part of the Dna.[73] Researchers from the University of Cambridge and the University of Illinois at Urbana-Champaign then demonstrated that such a DNA-induced toroidal pore tin facilitate rapid lipid flip-bomb between the lipid bilayer leaflets. Utilizing this event, they designed a synthetic DNA-built enzyme that flips lipids in biological membranes orders of magnitudes faster than naturally occurring proteins called scramblases.[80] This development highlights the potential of synthetic Dna nanostructures for personalized drugs and therapeutics.
Design [edit]
Dna nanostructures must be rationally designed so that individual nucleic acid strands will assemble into the desired structures. This procedure usually begins with specification of a desired target construction or function. Then, the overall secondary structure of the target complex is adamant, specifying the arrangement of nucleic acid strands within the structure, and which portions of those strands should be jump to each other. The last footstep is the principal structure design, which is the specification of the actual base sequences of each nucleic acid strand.[23] [81]
Structural pattern [edit]
The commencement step in designing a nucleic acid nanostructure is to make up one's mind how a given structure should exist represented by a specific organization of nucleic acid strands. This design step determines the secondary structure, or the positions of the base pairs that hold the private strands together in the desired shape.[23] Several approaches accept been demonstrated:
- Tile-based structures. This approach breaks the target construction into smaller units with strong binding between the strands contained in each unit of measurement, and weaker interactions between the units. It is often used to brand periodic lattices, but tin can too exist used to implement algorithmic self-assembly, making them a platform for DNA computing. This was the dominant design strategy used from the mid-1990s until the mid-2000s, when the Dna origami methodology was developed.[23] [82]
- Folding structures. An culling to the tile-based arroyo, folding approaches make the nanostructure from one long strand, which can either have a designed sequence that folds due to its interactions with itself, or information technology can be folded into the desired shape past using shorter, "staple" strands. This latter method is called DNA origami, which allows forming nanoscale two- and 3-dimensional shapes (encounter Discrete structures above).[27] [30]
- Dynamic assembly. This approach directly controls the kinetics of DNA self-assembly, specifying all of the intermediate steps in the reaction mechanism in addition to the concluding product. This is done using starting materials which adopt a hairpin structure; these then assemble into the concluding conformation in a cascade reaction, in a specific order (see Strand displacement cascades below). This approach has the advantage of proceeding isothermally, at a constant temperature. This is in contrast to the thermodynamic approaches, which require a thermal annealing step where a temperature modify is required to trigger the assembly and favor proper formation of the desired structure.[27] [58]
Sequence pattern [edit]
After any of the above approaches are used to design the secondary structure of a target circuitous, an actual sequence of nucleotides that will form into the desired construction must be devised. Nucleic acid design is the procedure of assigning a specific nucleic acrid base sequence to each of a construction'south constituent strands and so that they will associate into a desired conformation. Almost methods have the goal of designing sequences and then that the target structure has the lowest energy, and is thus the most thermodynamically favorable, while incorrectly assembled structures have college energies and are thus disfavored. This is washed either through simple, faster heuristic methods such as sequence symmetry minimization, or by using a full nearest-neighbor thermodynamic model, which is more than accurate only slower and more computationally intensive. Geometric models are used to examine third construction of the nanostructures and to ensure that the complexes are not overly strained.[81] [83]
Nucleic acid pattern has similar goals to protein blueprint. In both, the sequence of monomers is designed to favor the desired target structure and to disfavor other structures. Nucleic acid design has the reward of being much computationally easier than protein design, because the elementary base pairing rules are sufficient to predict a structure'due south energetic favorability, and detailed information about the overall three-dimensional folding of the structure is not required. This allows the use of simple heuristic methods that yield experimentally robust designs. Nucleic acid structures are less versatile than proteins in their function considering of proteins' increased power to fold into complex structures, and the express chemical diverseness of the four nucleotides as compared to the xx proteinogenic amino acids.[83]
Materials and methods [edit]
The sequences of the DNA strands making up a target construction are designed computationally, using molecular modeling and thermodynamic modeling software.[81] [83] The nucleic acids themselves are then synthesized using standard oligonucleotide synthesis methods, usually automated in an oligonucleotide synthesizer, and strands of custom sequences are commercially bachelor.[84] Strands can be purified past denaturing gel electrophoresis if needed,[85] and precise concentrations adamant via whatsoever of several nucleic acid quantitation methods using ultraviolet absorbance spectroscopy.[86]
The fully formed target structures tin be verified using native gel electrophoresis, which gives size and shape data for the nucleic acid complexes. An electrophoretic mobility shift assay tin assess whether a construction incorporates all desired strands.[87] Fluorescent labeling and Förster resonance energy transfer (FRET) are sometimes used to characterize the structure of the complexes.[88]
Nucleic acid structures can be direct imaged past diminutive forcefulness microscopy, which is well suited to extended 2-dimensional structures, but less useful for discrete iii-dimensional structures because of the microscope tip's interaction with the frail nucleic acid structure; transmission electron microscopy and cryo-electron microscopy are ofttimes used in this example. Extended three-dimensional lattices are analyzed by X-ray crystallography.[89] [90]
History [edit]

The woodcut Depth (pictured) by M. C. Escher reportedly inspired Nadrian Seeman to consider using 3-dimensional lattices of Dna to orient hard-to-crystallize molecules. This led to the get-go of the field of DNA nanotechnology.
The conceptual foundation for Deoxyribonucleic acid nanotechnology was kickoff laid out by Nadrian Seeman in the early 1980s.[91] Seeman's original motivation was to create a three-dimensional DNA lattice for orienting other large molecules, which would simplify their crystallographic study by eliminating the difficult process of obtaining pure crystals. This idea had reportedly come up to him in belatedly 1980, after realizing the similarity betwixt the woodcut Depth by Chiliad. C. Escher and an array of Dna 6-arm junctions.[3] [92] Several natural branched Deoxyribonucleic acid structures were known at the time, including the DNA replication fork and the mobile Holliday junction, but Seeman'due south insight was that immobile nucleic acid junctions could be created by properly designing the strand sequences to remove symmetry in the assembled molecule, and that these immobile junctions could in principle be combined into rigid crystalline lattices. The showtime theoretical newspaper proposing this scheme was published in 1982, and the first experimental demonstration of an immobile DNA junction was published the post-obit year.[v] [27]
In 1991, Seeman's laboratory published a written report on the synthesis of a cube fabricated of Dna, the first synthetic three-dimensional nucleic acid nanostructure, for which he received the 1995 Feynman Prize in Nanotechnology. This was followed by a DNA truncated octahedron. Information technology shortly became articulate that these structures, polygonal shapes with flexible junctions as their vertices, were non rigid enough to form extended iii-dimensional lattices. Seeman adult the more rigid double-crossover (DX) structural motif, and in 1998, in collaboration with Erik Winfree, published the creation of ii-dimensional lattices of DX tiles.[3] [91] [93] These tile-based structures had the advantage that they provided the ability to implement DNA computing, which was demonstrated by Winfree and Paul Rothemund in their 2004 paper on the algorithmic self-assembly of a Sierpinski gasket construction, and for which they shared the 2006 Feynman Prize in Nanotechnology. Winfree'south fundamental insight was that the DX tiles could exist used as Wang tiles, significant that their assembly could perform ciphering.[91] The synthesis of a three-dimensional lattice was finally published past Seeman in 2009, about thirty years after he had gear up out to accomplish it.[62]
New abilities continued to be discovered for designed DNA structures throughout the 2000s. The commencement Dna nanomachine—a motif that changes its structure in response to an input—was demonstrated in 1999 past Seeman. An improved system, which was the start nucleic acid device to make use of toehold-mediated strand deportation, was demonstrated past Bernard Yurke the post-obit twelvemonth. The next advance was to translate this into mechanical move, and in 2004 and 2005, several DNA walker systems were demonstrated past the groups of Seeman, Niles Pierce, Andrew Turberfield, and Chengde Mao.[42] The idea of using DNA arrays to template the associates of other molecules such as nanoparticles and proteins, first suggested past Bruche Robinson and Seeman in 1987,[94] was demonstrated in 2002 by Seeman, Kiehl et al.[95] and subsequently by many other groups.
In 2006, Rothemund first demonstrated the DNA origami method for easily and robustly forming folded Deoxyribonucleic acid structures of arbitrary shape. Rothemund had conceived of this method equally being conceptually intermediate between Seeman's DX lattices, which used many short strands, and William Shih's Dna octahedron, which consisted mostly of ane very long strand. Rothemund's Deoxyribonucleic acid origami contains a long strand which folding is assisted by several short strands. This method allowed forming much larger structures than formerly possible, and which are less technically enervating to blueprint and synthesize.[93] DNA origami was the cover story of Nature on March 15, 2006.[30] Rothemund's research demonstrating 2-dimensional DNA origami structures was followed past the demonstration of solid iii-dimensional Deoxyribonucleic acid origami by Douglas et al. in 2009,[32] while the labs of Jørgen Kjems and Yan demonstrated hollow three-dimensional structures fabricated out of ii-dimensional faces.[62]
DNA nanotechnology was initially met with some skepticism due to the unusual not-biological use of nucleic acids as materials for edifice structures and doing ciphering, and the preponderance of proof of principle experiments that extended the abilities of the field but were far from actual applications. Seeman's 1991 paper on the synthesis of the DNA cube was rejected by the journal Scientific discipline afterwards i reviewer praised its originality while another criticized it for its lack of biological relevance.[96] By the early 2010s the field was considered to accept increased its abilities to the indicate that applications for bones scientific discipline enquiry were offset to exist realized, and practical applications in medicine and other fields were beginning to exist considered viable.[62] [97] The field had grown from very few active laboratories in 2001 to at least 60 in 2010, which increased the talent pool and thus the number of scientific advances in the field during that decade.[21]
See likewise [edit]
- International Society for Nanoscale Science, Computation, and Technology
- Comparison of nucleic acid simulation software
- Molecular models of DNA
- Nanobiotechnology
References [edit]
- ^ a b Dna polyhedra: Goodman RP, Schaap IA, Tardin CF, Erben CM, Berry RM, Schmidt CF, Turberfield AJ (December 2005). "Rapid chiral associates of rigid Deoxyribonucleic acid edifice blocks for molecular nanofabrication". Science. 310 (5754): 1661–1665. Bibcode:2005Sci...310.1661G. doi:10.1126/science.1120367. PMID 16339440. S2CID 13678773.
- ^ a b c Overview: Mao C (December 2004). "The emergence of complication: lessons from DNA". PLOS Biology. two (12): e431. doi:10.1371/journal.pbio.0020431. PMC535573. PMID 15597116.
- ^ a b c d e Overview: Seeman NC (June 2004). "Nanotechnology and the double helix". Scientific American. 290 (6): 64–75. Bibcode:2004SciAm.290f..64S. doi:10.1038/scientificamerican0604-64. PMID 15195395.
- ^ Background: Pelesko JA (2007). Self-assembly: the science of things that put themselves together. New York: Chapman & Hall/CRC. pp. five, 7. ISBN978-1-58488-687-vii.
- ^ a b c d e Overview: Seeman NC (2010). "Nanomaterials based on Dna". Annual Review of Biochemistry. 79: 65–87. doi:ten.1146/annurev-biochem-060308-102244. PMC3454582. PMID 20222824.
- ^ Groundwork: Long EC (1996). "Fundamentals of nucleic acids". In Hecht SM (ed.). Bioorganic chemical science: nucleic acids. New York: Oxford Academy Printing. pp. 4–10. ISBN978-0-19-508467-two.
- ^ RNA nanotechnology: Chworos A, Severcan I, Koyfman AY, Weinkam P, Oroudjev East, Hansma HG, Jaeger L (December 2004). "Building programmable jigsaw puzzles with RNA". Science. 306 (5704): 2068–2072. Bibcode:2004Sci...306.2068C. doi:10.1126/science.1104686. PMID 15604402. S2CID 9296608.
- ^ RNA nanotechnology: Guo P (December 2010). "The emerging field of RNA nanotechnology". Nature Nanotechnology. v (12): 833–842. Bibcode:2010NatNa...5..833G. doi:ten.1038/nnano.2010.231. PMC3149862. PMID 21102465.
- ^ a b c d Dynamic Dna nanotechnology: Zhang DY, Seelig G (February 2011). "Dynamic DNA nanotechnology using strand-displacement reactions". Nature Chemical science. 3 (2): 103–113. Bibcode:2011NatCh...3..103Z. doi:x.1038/nchem.957. PMID 21258382.
- ^ a b c d e Structural Dna nanotechnology: Seeman NC (Nov 2007). "An overview of structural DNA nanotechnology". Molecular Biotechnology. 37 (iii): 246–257. doi:10.1007/s12033-007-0059-four. PMC3479651. PMID 17952671.
- ^ Dynamic Dna nanotechnology: Lu Y, Liu J (December 2006). "Functional DNA nanotechnology: emerging applications of DNAzymes and aptamers". Current Opinion in Biotechnology. 17 (6): 580–588. doi:10.1016/j.copbio.2006.10.004. PMID 17056247.
- ^ Simulation of DNA structures: Doye JP, Ouldridge TE, Louis AA, Romano F, Šulc P, Matek C, et al. (December 2013). "Coarse-graining Dna for simulations of DNA nanotechnology". Concrete Chemistry Chemic Physics. fifteen (47): 20395–20414. arXiv:1308.3843. Bibcode:2013PCCP...1520395D. doi:10.1039/C3CP53545B. PMID 24121860. S2CID 15324396.
- ^ Other arrays: Strong M (March 2004). "Protein nanomachines". PLOS Biology. ii (3): E73. doi:ten.1371/journal.pbio.0020073. PMC368168. PMID 15024422.
- ^ Yan H, Park SH, Finkelstein G, Reif JH, LaBean TH (September 2003). "DNA-templated self-assembly of protein arrays and highly conductive nanowires". Scientific discipline. 301 (5641): 1882–1884. Bibcode:2003Sci...301.1882Y. doi:10.1126/science.1089389. PMID 14512621. S2CID 137635908.
- ^ a b Algorithmic self-assembly: Rothemund Pw, Papadakis N, Winfree E (December 2004). "Algorithmic cocky-assembly of DNA Sierpinski triangles". PLOS Biology. two (12): e424. doi:ten.1371/journal.pbio.0020424. PMC534809. PMID 15583715.
- ^ DX arrays: Winfree East, Liu F, Wenzler LA, Seeman NC (August 1998). "Blueprint and self-assembly of 2-dimensional DNA crystals". Nature. 394 (6693): 539–544. Bibcode:1998Natur.394..539W. doi:10.1038/28998. PMID 9707114. S2CID 4385579.
- ^ DX arrays: Liu F, Sha R, Seeman NC (10 Feb 1999). "Modifying the surface features of two-dimensional Deoxyribonucleic acid crystals". Journal of the American Chemical Guild. 121 (5): 917–922. doi:ten.1021/ja982824a.
- ^ Other arrays: Mao C, Sun W, Seeman NC (xvi June 1999). "Designed two-dimensional Dna Holliday junction arrays visualized by atomic forcefulness microscopy". Journal of the American Chemical Society. 121 (23): 5437–5443. doi:x.1021/ja9900398.
- ^ Other arrays: Constantinou PE, Wang T, Kopatsch J, Israel LB, Zhang X, Ding B, et al. (September 2006). "Double cohesion in structural Deoxyribonucleic acid nanotechnology". Organic & Biomolecular Chemical science. iv (18): 3414–3419. doi:ten.1039/b605212f. PMC3491902. PMID 17036134.
- ^ Other arrays: Mathieu F, Liao Southward, Kopatsch J, Wang T, Mao C, Seeman NC (April 2005). "Six-helix bundles designed from DNA". Nano Messages. 5 (4): 661–665. Bibcode:2005NanoL...v..661M. doi:10.1021/nl050084f. PMC3464188. PMID 15826105.
- ^ a b c History: Seeman NC (June 2010). "Structural DNA nanotechnology: growing forth with Nano Messages". Nano Letters. 10 (half dozen): 1971–1978. Bibcode:2010NanoL..10.1971S. doi:10.1021/nl101262u. PMC2901229. PMID 20486672.
- ^ Algorithmic self-assembly: Barish RD, Rothemund Prisoner of war, Winfree E (December 2005). "Two computational primitives for algorithmic self-assembly: copying and counting". Nano Messages. 5 (12): 2586–2592. Bibcode:2005NanoL...v.2586B. CiteSeerX10.1.i.155.676. doi:10.1021/nl052038l. PMID 16351220.
- ^ a b c d Blueprint: Feldkamp U, Niemeyer CM (March 2006). "Rational blueprint of Dna nanoarchitectures". Angewandte Chemie. 45 (12): 1856–1876. doi:10.1002/anie.200502358. PMID 16470892.
- ^ DNA nanotubes: Rothemund Pow, Ekani-Nkodo A, Papadakis N, Kumar A, Fygenson DK, Winfree Due east (December 2004). "Blueprint and characterization of programmable DNA nanotubes". Journal of the American Chemical Society. 126 (50): 16344–16352. doi:10.1021/ja044319l. PMID 15600335.
- ^ DNA nanotubes: Yin P, Hariadi RF, Sahu S, Choi HM, Park SH, Labean TH, Reif JH (Baronial 2008). "Programming DNA tube circumferences". Science. 321 (5890): 824–826. Bibcode:2008Sci...321..824Y. doi:10.1126/science.1157312. PMID 18687961. S2CID 12100380.
- ^ Three-dimensional arrays: Zheng J, Birktoft JJ, Chen Y, Wang T, Sha R, Constantinou PE, et al. (September 2009). "From molecular to macroscopic via the rational blueprint of a self-assembled 3D DNA crystal". Nature. 461 (7260): 74–77. Bibcode:2009Natur.461...74Z. doi:x.1038/nature08274. PMC2764300. PMID 19727196.
- ^ a b c d e f g h i Overview: Pinheiro AV, Han D, Shih WM, Yan H (Nov 2011). "Challenges and opportunities for structural DNA nanotechnology". Nature Nanotechnology. 6 (12): 763–772. Bibcode:2011NatNa...six..763P. doi:10.1038/nnano.2011.187. PMC3334823. PMID 22056726.
- ^ DNA polyhedra: Zhang Y, Seeman NC (1 March 1994). "Construction of a DNA-truncated octahedron". Journal of the American Chemical Social club. 116 (5): 1661–1669. doi:ten.1021/ja00084a006.
- ^ Deoxyribonucleic acid polyhedra: Shih WM, Quispe JD, Joyce GF (February 2004). "A i.seven-kilobase single-stranded DNA that folds into a nanoscale octahedron". Nature. 427 (6975): 618–621. Bibcode:2004Natur.427..618S. doi:10.1038/nature02307. PMID 14961116. S2CID 4419579.
- ^ a b c DNA origami: Rothemund PW (March 2006). "Folding Deoxyribonucleic acid to create nanoscale shapes and patterns" (PDF). Nature. 440 (7082): 297–302. Bibcode:2006Natur.440..297R. doi:ten.1038/nature04586. PMID 16541064. S2CID 4316391.
- ^ Tikhomirov G, Petersen P, Qian L (December 2017). "Fractal assembly of micrometre-scale DNA origami arrays with capricious patterns". Nature. 552 (7683): 67–71. Bibcode:2017Natur.552...67T. doi:10.1038/nature24655. PMID 29219965. S2CID 4455780.
- ^ a b Dna origami: Douglas SM, Dietz H, Liedl T, Högberg B, Graf F, Shih WM (May 2009). "Self-assembly of DNA into nanoscale three-dimensional shapes". Nature. 459 (7245): 414–418. Bibcode:2009Natur.459..414D. doi:10.1038/nature08016. PMC2688462. PMID 19458720.
- ^ a b Dna boxes: Andersen ES, Dong K, Nielsen MM, Jahn K, Subramani R, Mamdouh W, et al. (May 2009). "Self-assembly of a nanoscale DNA box with a controllable lid". Nature. 459 (7243): 73–76. Bibcode:2009Natur.459...73A. doi:10.1038/nature07971. hdl:11858/00-001M-0000-0010-9363-9. PMID 19424153. S2CID 4430815.
- ^ DNA boxes: Ke Y, Sharma J, Liu Thousand, Jahn K, Liu Y, Yan H (June 2009). "Scaffolded Dna origami of a Dna tetrahedron molecular container". Nano Letters. nine (half-dozen): 2445–2447. Bibcode:2009NanoL...9.2445K. doi:ten.1021/nl901165f. PMID 19419184.
- ^ Overview: Endo M, Sugiyama H (October 2009). "Chemical approaches to Deoxyribonucleic acid nanotechnology". ChemBioChem. 10 (15): 2420–2443. doi:10.1002/cbic.200900286. PMID 19714700. S2CID 205554125.
- ^ Nanoarchitecture: Zheng J, Constantinou PE, Micheel C, Alivisatos AP, Kiehl RA, Seeman NC (July 2006). "Two-dimensional nanoparticle arrays testify the organizational power of robust DNA motifs". Nano Letters. 6 (7): 1502–1504. Bibcode:2006NanoL...six.1502Z. doi:x.1021/nl060994c. PMC3465979. PMID 16834438.
- ^ Nanoarchitecture: Park SH, Pistol C, Ahn SJ, Reif JH, Lebeck AR, Dwyer C, LaBean Thursday (January 2006). "Finite-size, fully addressable DNA tile lattices formed by hierarchical associates procedures". Angewandte Chemie. 45 (v): 735–739. Bibcode:2006AngCh.118.6759P. doi:10.1002/ange.200690141. PMID 16374784.
- ^ Nanoarchitecture: Cohen JD, Sadowski JP, Dervan Atomic number 82 (22 October 2007). "Addressing single molecules on DNA nanostructures". Angewandte Chemie. 46 (42): 7956–7959. doi:10.1002/anie.200702767. PMID 17763481.
- ^ Nanoarchitecture: Maune HT, Han SP, Barish RD, Bockrath M, Goddard WA, Rothemund PW, Winfree E (Jan 2010). "Self-associates of carbon nanotubes into two-dimensional geometries using Deoxyribonucleic acid origami templates". Nature Nanotechnology. 5 (1): 61–66. Bibcode:2010NatNa...5...61M. doi:10.1038/nnano.2009.311. PMID 19898497.
- ^ Nanoarchitecture: Liu J, Geng Y, Pound East, Gyawali South, Ashton JR, Hickey J, et al. (March 2011). "Metallization of branched Deoxyribonucleic acid origami for nanoelectronic circuit fabrication". ACS Nano. 5 (3): 2240–2247. doi:10.1021/nn1035075. PMID 21323323.
- ^ Nanoarchitecture: Deng Z, Mao C (August 2004). "Molecular lithography with Deoxyribonucleic acid nanostructures". Angewandte Chemie. 43 (31): 4068–4070. doi:10.1002/anie.200460257. PMID 15300697.
- ^ a b c d DNA machines: Bathroom J, Turberfield AJ (May 2007). "DNA nanomachines". Nature Nanotechnology. two (5): 275–284. Bibcode:2007NatNa...2..275B. doi:10.1038/nnano.2007.104. PMID 18654284.
- ^ DNA machines: Mao C, Sunday W, Shen Z, Seeman NC (January 1999). "A nanomechanical device based on the B-Z transition of DNA". Nature. 397 (6715): 144–146. Bibcode:1999Natur.397..144M. doi:ten.1038/16437. PMID 9923675. S2CID 4406177.
- ^ DNA machines: Yurke B, Turberfield AJ, Mills AP, Simmel FC, Neumann JL (August 2000). "A DNA-fuelled molecular machine made of DNA". Nature. 406 (6796): 605–608. Bibcode:2000Natur.406..605Y. doi:x.1038/35020524. PMID 10949296. S2CID 2064216.
- ^ DNA machines: Yan H, Zhang X, Shen Z, Seeman NC (January 2002). "A robust Deoxyribonucleic acid mechanical device controlled by hybridization topology". Nature. 415 (6867): 62–65. Bibcode:2002Natur.415...62Y. doi:x.1038/415062a. PMID 11780115. S2CID 52801697.
- ^ DNA machines: Feng L, Park SH, Reif JH, Yan H (September 2003). "A ii-state Dna lattice switched by Deoxyribonucleic acid nanoactuator". Angewandte Chemie. 42 (36): 4342–4346. Bibcode:2003AngCh.115.4478F. doi:ten.1002/ange.200351818. PMID 14502706.
- ^ Deoxyribonucleic acid machines: Goodman RP, Heilemann M, Doose S, Erben CM, Kapanidis AN, Turberfield AJ (February 2008). "Reconfigurable, braced, three-dimensional Deoxyribonucleic acid nanostructures". Nature Nanotechnology. 3 (2): 93–96. Bibcode:2008NatNa...3...93G. doi:10.1038/nnano.2008.3. PMID 18654468.
- ^ Applications: Douglas SM, Bachelet I, Church GM (February 2012). "A logic-gated nanorobot for targeted transport of molecular payloads". Science. 335 (6070): 831–834. Bibcode:2012Sci...335..831D. doi:10.1126/science.1214081. PMID 22344439. S2CID 9866509.
- ^ Deoxyribonucleic acid walkers: Shin JS, Pierce NA (September 2004). "A constructed DNA walker for molecular transport". Periodical of the American Chemical Gild. 126 (35): 10834–10835. doi:x.1021/ja047543j. PMID 15339155.
- ^ Deoxyribonucleic acid walkers: Sherman WB, Seeman NC (July 2004). "A precisely controlled Dna biped walking device". Nano Letters. 4 (vii): 1203–1207. Bibcode:2004NanoL...four.1203S. doi:x.1021/nl049527q.
- ^ DNA walkers: Škugor K, Valero J, Murayama M, Centola M, Asanuma H, Famulok 1000 (May 2019). "Orthogonally Photocontrolled Non-Autonomous DNA Walker". Angewandte Chemie. 58 (21): 6948–6951. doi:10.1002/anie.201901272. PMID 30897257. S2CID 85446523.
- ^ Deoxyribonucleic acid walkers: Tian Y, He Y, Chen Y, Yin P, Mao C (July 2005). "A DNAzyme that walks processively and autonomously along a one-dimensional rails". Angewandte Chemie. 44 (28): 4355–4358. Bibcode:2005AngCh.117.4429T. doi:10.1002/ange.200500703. PMID 15945114.
- ^ DNA walkers: Bath J, Green SJ, Turberfield AJ (July 2005). "A complimentary-running Deoxyribonucleic acid motor powered past a nicking enzyme". Angewandte Chemie. 44 (28): 4358–4361. doi:10.1002/anie.200501262. PMID 15959864.
- ^ Functional Deoxyribonucleic acid walkers: Lund 1000, Manzo AJ, Dabby N, Michelotti N, Johnson-Buck A, Nangreave J, et al. (May 2010). "Molecular robots guided past prescriptive landscapes". Nature. 465 (7295): 206–210. Bibcode:2010Natur.465..206L. doi:x.1038/nature09012. PMC2907518. PMID 20463735.
- ^ Functional Dna walkers: Valero J, Pal N, Dhakal South, Walter NG, Famulok M (June 2018). "A bio-hybrid DNA rotor-stator nanoengine that moves along predefined tracks". Nature Nanotechnology. xiii (half-dozen): 496–503. Bibcode:2018NatNa..13..496V. doi:10.1038/s41565-018-0109-z. PMC5994166. PMID 29632399.
- ^ Functional DNA walkers: He Y, Liu DR (Nov 2010). "Autonomous multistep organic synthesis in a unmarried isothermal solution mediated by a DNA walker". Nature Nanotechnology. 5 (xi): 778–782. Bibcode:2010NatNa...5..778H. doi:10.1038/nnano.2010.190. PMC2974042. PMID 20935654.
- ^ Pan J, Li F, Cha TG, Chen H, Choi JH (August 2015). "Recent progress on Deoxyribonucleic acid based walkers". Current Opinion in Biotechnology. 34: 56–64. doi:ten.1016/j.copbio.2014.xi.017. PMID 25498478.
- ^ a b c Kinetic assembly: Yin P, Choi HM, Calvert CR, Pierce NA (January 2008). "Programming biomolecular self-assembly pathways". Nature. 451 (7176): 318–322. Bibcode:2008Natur.451..318Y. doi:x.1038/nature06451. PMID 18202654. S2CID 4354536.
- ^ Fuzzy and Boolean logic gates based on DNA: Zadegan RM, Jepsen Dr., Hildebrandt LL, Birkedal V, Kjems J (Apr 2015). "Structure of a fuzzy and Boolean logic gates based on DNA". Small. 11 (15): 1811–1817. doi:10.1002/smll.201402755. PMID 25565140.
- ^ Strand displacement cascades: Seelig G, Soloveichik D, Zhang DY, Winfree E (December 2006). "Enzyme-gratuitous nucleic acid logic circuits". Science. 314 (5805): 1585–1588. Bibcode:2006Sci...314.1585S. doi:10.1126/science.1132493. PMID 17158324. S2CID 10966324.
- ^ Strand displacement cascades: Qian L, Winfree Due east (June 2011). "Scaling up digital circuit computation with Dna strand displacement cascades". Science. 332 (6034): 1196–1201. Bibcode:2011Sci...332.1196Q. doi:10.1126/science.1200520. PMID 21636773. S2CID 10053541.
- ^ a b c d e History/applications: Service RF (June 2011). "Dna nanotechnology. Deoxyribonucleic acid nanotechnology grows up". Scientific discipline. 332 (6034): 1140–1, 1143. Bibcode:2011Sci...332.1140S. doi:ten.1126/science.332.6034.1140. PMID 21636754.
- ^ Applications: Rietman EA (2001). Molecular engineering of nanosystems. Springer. pp. 209–212. ISBN978-0-387-98988-4 . Retrieved 17 Apr 2011.
- ^ Zadegan RM, Jepsen MD, Thomsen KE, Okholm AH, Schaffert DH, Andersen ES, et al. (November 2012). "Construction of a 4 zeptoliters switchable 3D Dna box origami". ACS Nano. 6 (xi): 10050–10053. doi:x.1021/nn303767b. PMID 23030709.
- ^ Applications: Jungmann R, Renner S, Simmel FC (April 2008). "From Dna nanotechnology to synthetic biology". HFSP Journal. ii (two): 99–109. doi:10.2976/ane.2896331. PMC2645571. PMID 19404476.
- ^ Lovy, Howard (5 July 2011). "DNA cages can unleash meds inside cells". fiercedrugdelivery.com. Retrieved 22 September 2013.
- ^ Walsh Equally, Yin H, Erben CM, Forest MJ, Turberfield AJ (July 2011). "DNA muzzle commitment to mammalian cells". ACS Nano. 5 (seven): 5427–5432. doi:10.1021/nn2005574. PMID 21696187.
- ^ Trafton, Anne (4 June 2012). "Researchers achieve RNA interference, in a lighter packet". MIT News. Retrieved 22 September 2013.
- ^ Lee H, Lytton-Jean AK, Chen Y, Love KT, Park AI, Karagiannis ED, et al. (June 2012). "Molecularly self-assembled nucleic acid nanoparticles for targeted in vivo siRNA delivery". Nature Nanotechnology. 7 (6): 389–393. Bibcode:2012NatNa...7..389L. doi:10.1038/NNANO.2012.73. PMC3898745. PMID 22659608.
- ^ Kim KR, Kim DR, Lee T, Yhee JY, Kim BS, Kwon IC, Ahn DR (March 2013). "Drug delivery by a self-assembled Dna tetrahedron for overcoming drug resistance in chest cancer cells". Chemic Communications. 49 (20): 2010–2012. doi:10.1039/c3cc38693g. PMID 23380739.
- ^ Sundah NR, Ho NR, Lim GS, Natalia A, Ding X, Liu Y, et al. (September 2019). "Barcoded Deoxyribonucleic acid nanostructures for the multiplexed profiling of subcellular protein distribution". Nature Biomedical Technology. 3 (9): 684–694. doi:10.1038/s41551-019-0417-0. PMID 31285580. S2CID 195825879.
- ^ Dna ion channels: Langecker M, Arnaut V, Martin TG, List J, Renner Southward, Mayer Yard, et al. (November 2012). "Constructed lipid membrane channels formed by designed DNA nanostructures". Science. 338 (6109): 932–936. Bibcode:2012Sci...338..932L. doi:10.1126/science.1225624. PMC3716461. PMID 23161995.
- ^ a b Dna ion channels: Göpfrich K, Li CY, Mames I, Bhamidimarri SP, Ricci M, Yoo J, et al. (July 2016). "Ion Channels Fabricated from a Single Membrane-Spanning Deoxyribonucleic acid Duplex". Nano Letters. 16 (7): 4665–4669. Bibcode:2016NanoL..xvi.4665G. doi:ten.1021/acs.nanolett.6b02039. PMC4948918. PMID 27324157.
- ^ DNA ion channels: Burns JR, Stulz E, Howorka S (June 2013). "Cocky-assembled DNA nanopores that bridge lipid bilayers". Nano Letters. thirteen (vi): 2351–2356. Bibcode:2013NanoL..13.2351B. CiteSeerXten.1.one.659.7660. doi:10.1021/nl304147f. PMID 23611515.
- ^ Deoxyribonucleic acid ion channels: Burns JR, Göpfrich Thousand, Wood JW, Thacker VV, Stulz E, Keyser UF, Howorka S (November 2013). "Lipid-bilayer-spanning DNA nanopores with a bifunctional porphyrin anchor". Angewandte Chemie. 52 (46): 12069–12072. doi:x.1002/anie.201305765. PMC4016739. PMID 24014236.
- ^ Deoxyribonucleic acid ion channels: Seifert A, Göpfrich G, Burns JR, Fertig N, Keyser UF, Howorka S (February 2015). "Bilayer-spanning DNA nanopores with voltage-switching between open and closed state". ACS Nano. 9 (2): 1117–1126. doi:10.1021/nn5039433. PMC4508203. PMID 25338165.
- ^ Dna ion channels: Göpfrich 1000, Zettl T, Meijering AE, Hernández-Ainsa South, Kocabey Southward, Liedl T, Keyser UF (May 2015). "Dna-Tile Structures Induce Ionic Currents through Lipid Membranes". Nano Letters. 15 (5): 3134–3138. Bibcode:2015NanoL..15.3134G. doi:10.1021/acs.nanolett.5b00189. PMID 25816075.
- ^ Dna ion channels: Burns JR, Seifert A, Fertig N, Howorka Due south (February 2016). "A biomimetic Deoxyribonucleic acid-based channel for the ligand-controlled ship of charged molecular cargo across a biological membrane". Nature Nanotechnology. eleven (two): 152–156. Bibcode:2016NatNa..11..152B. doi:10.1038/nnano.2015.279. PMID 26751170.
- ^ DNA ion channels: Göpfrich K, Li CY, Ricci M, Bhamidimarri SP, Yoo J, Gyenes B, et al. (September 2016). "Large-Conductance Transmembrane Porin Made from DNA Origami". ACS Nano. 10 (nine): 8207–8214. doi:10.1021/acsnano.6b03759. PMC5043419. PMID 27504755.
- ^ Dna scramblase: Ohmann A, Li CY, Maffeo C, Al Nahas G, Baumann KN, Göpfrich K, et al. (June 2018). "A synthetic enzyme built from Deoxyribonucleic acid flips 107 lipids per 2d in biological membranes". Nature Communications. ix (1): 2426. Bibcode:2018NatCo...ix.2426O. doi:10.1038/s41467-018-04821-v. PMC6013447. PMID 29930243.
- ^ a b c Design: Brenneman A, Condon A (25 September 2002). "Strand design for biomolecular computation". Theoretical Computer Science. 287: 39–58. doi:10.1016/S0304-3975(02)00135-iv.
- ^ Overview: Lin C, Liu Y, Rinker Due south, Yan H (August 2006). "Deoxyribonucleic acid tile based self-assembly: building complex nanoarchitectures". ChemPhysChem. 7 (8): 1641–1647. doi:10.1002/cphc.200600260. PMID 16832805.
- ^ a b c Design: Dirks RM, Lin One thousand, Winfree Eastward, Pierce NA (15 Feb 2004). "Paradigms for computational nucleic acrid design". Nucleic Acids Research. 32 (4): 1392–1403. doi:10.1093/nar/gkh291. PMC390280. PMID 14990744.
- ^ Methods: Ellington A, Pollard JD (1 May 2001). Synthesis and Purification of Oligonucleotides. Current Protocols in Molecular Biology. Vol. Chapter 2. pp. 2.eleven.ane–two.11.25. doi:x.1002/0471142727.mb0211s42. ISBN978-0471142720. PMID 18265179. S2CID 205152989.
- ^ Methods: Ellington A, Pollard JD (i May 2001). Purification of Oligonucleotides Using Denaturing Polyacrylamide Gel Electrophoresis. Current Protocols in Molecular Biology. Vol. Affiliate two. pp. Unit2.12. doi:ten.1002/0471142727.mb0212s42. ISBN978-0471142720. PMID 18265180. S2CID 27187583.
- ^ Methods: Gallagher SR, Desjardins P (ane July 2011). "Quantitation of nucleic acids and proteins". Current Protocols Essential Laboratory techniques. Vol. five. doi:10.1002/9780470089941.et0202s5. ISBN978-0470089934. S2CID 94329398.
- ^ Methods: Chory J, Pollard JD (1 May 2001). Separation of Pocket-size Dna Fragments by Conventional Gel Electrophoresis. Current Protocols in Molecular Biology. Vol. Chapter ii. pp. Unit2.7. doi:10.1002/0471142727.mb0207s47. ISBN978-0471142720. PMID 18265187. S2CID 43406338.
- ^ Methods: Walter NG (i Feb 2003). "Probing RNA structural dynamics and function by fluorescence resonance energy transfer (FRET)". Electric current Protocols in Nucleic Acrid Chemistry. Current Protocols in Nucleic Acrid Chemistry. Vol. Chapter 11. pp. eleven.10.1–11.ten.23. doi:10.1002/0471142700.nc1110s11. ISBN978-0471142706. PMID 18428904.
- ^ Methods: Lin C, Ke Y, Chhabra R, Sharma J, Liu Y, Yan H (2011). "Synthesis and characterization of self-assembled DNA nanostructures". In Zuccheri G, Samorì B (eds.). Deoxyribonucleic acid Nanotechnology. Methods in Molecular Biology. Vol. 749. pp. 1–xi. doi:10.1007/978-1-61779-142-0_1. ISBN978-1-61779-141-3. PMID 21674361.
- ^ Methods: Bloomfield VA, Crothers DM, Tinoco Jr I (2000). Nucleic acids: structures, backdrop, and functions. Sausalito, Calif: University Science Books. pp. 84–86, 396–407. ISBN978-0-935702-49-ane.
- ^ a b c History: Pelesko JA (2007). Cocky-assembly: the science of things that put themselves together. New York: Chapman & Hall/CRC. pp. 201, 242, 259. ISBN978-1-58488-687-7.
- ^ History: See "Electric current crystallization protocol". Nadrian Seeman Lab. for a argument of the problem, and "DNA cages containing oriented guests". Nadrian Seeman Laboratory. for the proposed solution.
- ^ a b Dna origami: Rothemund Prisoner of war (2006). "Scaffolded Deoxyribonucleic acid origami: from generalized multicrossovers to polygonal networks". In Chen J, Jonoska Due north, Rozenberg Chiliad (eds.). Nanotechnology: scientific discipline and computation. Natural Computing Serial. New York: Springer. pp. 3–21. CiteSeerX10.1.1.144.1380. doi:x.1007/3-540-30296-4_1. ISBN978-three-540-30295-vii.
- ^ Nanoarchitecture: Robinson BH, Seeman NC (August 1987). "The design of a biochip: a self-assembling molecular-scale memory device". Poly peptide Engineering. i (4): 295–300. doi:10.1093/poly peptide/1.4.295. PMID 3508280.
- ^ Nanoarchitecture: Xiao Southward, Liu F, Rosen AE, Hainfeld JF, Seeman NC, Musier-Forsyth K, Kiehl RA (Baronial 2002). "Selfassembly of metal nanoparticle arrays by DNA scaffolding". Periodical of Nanoparticle Research. 4 (four): 313–317. Bibcode:2002JNR.....four..313X. doi:ten.1023/A:1021145208328. S2CID 2257083.
- ^ Service RF (June 2011). "Deoxyribonucleic acid nanotechnology. Dna nanotechnology grows up". Science. 332 (6034): 1140–one, 1143. Bibcode:2011Sci...332.1140S. doi:10.1126/scientific discipline.332.6034.1140. PMID 21636754.
- ^ History: Hopkin K (Baronial 2011). "Profile: 3-D seer". The Scientist. Archived from the original on ten Oct 2011. Retrieved 8 Baronial 2011.
Further reading [edit]
Full general:
- Seeman NC (June 2004). "Nanotechnology and the double helix". Scientific American. 290 (6): 64–75. Bibcode:2004SciAm.290f..64S. doi:10.1038/scientificamerican0604-64. PMID 15195395. —An article written for laypeople by the founder of the field
- Seeman NC (June 2010). "Structural DNA nanotechnology: growing along with Nano Messages". Nano Messages. ten (6): 1971–1978. Bibcode:2010NanoL..x.1971S. doi:10.1021/nl101262u. PMC2901229. PMID 20486672. —A review of results in the period 2001–2010
- Seeman NC (2010). "Nanomaterials based on Dna". Almanac Review of Biochemistry. 79: 65–87. doi:10.1146/annurev-biochem-060308-102244. PMC3454582. PMID 20222824. —A more comprehensive review including both old and new results in the field
- Service RF (June 2011). "DNA nanotechnology. Dna nanotechnology grows upward". Scientific discipline. 332 (6034): 1140–1, 1143. Bibcode:2011Sci...332.1140S. doi:10.1126/scientific discipline.332.6034.1140. PMID 21636754. and Service RF (June 2011). "DNA nanotechnology. Next footstep: Dna robots?". Science. 332 (6034): 1142. doi:10.1126/science.332.6034.1142. PMID 21636755. .—A news article focusing on the history of the field and development of new applications
- Zadegan RM, Norton ML (June 2012). "Structural Deoxyribonucleic acid nanotechnology: from design to applications". International Journal of Molecular Sciences. thirteen (6): 7149–7162. doi:10.3390/ijms13067149. PMC3397516. PMID 22837684. —A very recent and comprehensive review in the field
Specific subfields:
- Bath J, Turberfield AJ (May 2007). "DNA nanomachines". Nature Nanotechnology. 2 (v): 275–284. Bibcode:2007NatNa...2..275B. doi:x.1038/nnano.2007.104. PMID 18654284. —A review of nucleic acid nanomechanical devices
- Feldkamp U, Niemeyer CM (March 2006). "Rational design of DNA nanoarchitectures". Angewandte Chemie. 45 (12): 1856–1876. doi:x.1002/anie.200502358. PMID 16470892. —A review coming from the viewpoint of secondary structure design
- Lin C, Liu Y, Rinker Due south, Yan H (Baronial 2006). "Deoxyribonucleic acid tile based self-assembly: building complex nanoarchitectures". ChemPhysChem. 7 (8): 1641–1647. doi:ten.1002/cphc.200600260. PMID 16832805. —A minireview specifically focusing on tile-based assembly
- Zhang DY, Seelig Yard (February 2011). "Dynamic Deoxyribonucleic acid nanotechnology using strand-displacement reactions". Nature Chemical science. 3 (two): 103–113. Bibcode:2011NatCh...3..103Z. doi:10.1038/nchem.957. PMID 21258382. —A review of Dna systems making use of strand displacement mechanisms
External links [edit]
- What is Bionanotechnology?—a video introduction to Deoxyribonucleic acid nanotechnology
Source: https://en.wikipedia.org/wiki/DNA_nanotechnology
Post a Comment for "How to Read 2 Number Pairs Into an Array of Structures in C#"